MolCraftDiffusion¶
A unified generative-AI framework for 3D molecular design.
MolCraftDiffusion is an open-source framework for 3D molecular generation using diffusion models, designed for data-driven molecular design and computational chemistry. The framework enables researchers to train generative models that produce chemically meaningful 3D molecular structures while supporting property optimization, scaffold modification, and exploration of chemical space.
By combining modular training pipelines, flexible guidance strategies, and integrated analysis tools, MolCraftDiffusion provides a complete workflow for developing and deploying molecular diffusion models in research applications such as drug discovery, catalyst discovery, materials design, and molecular property optimization.
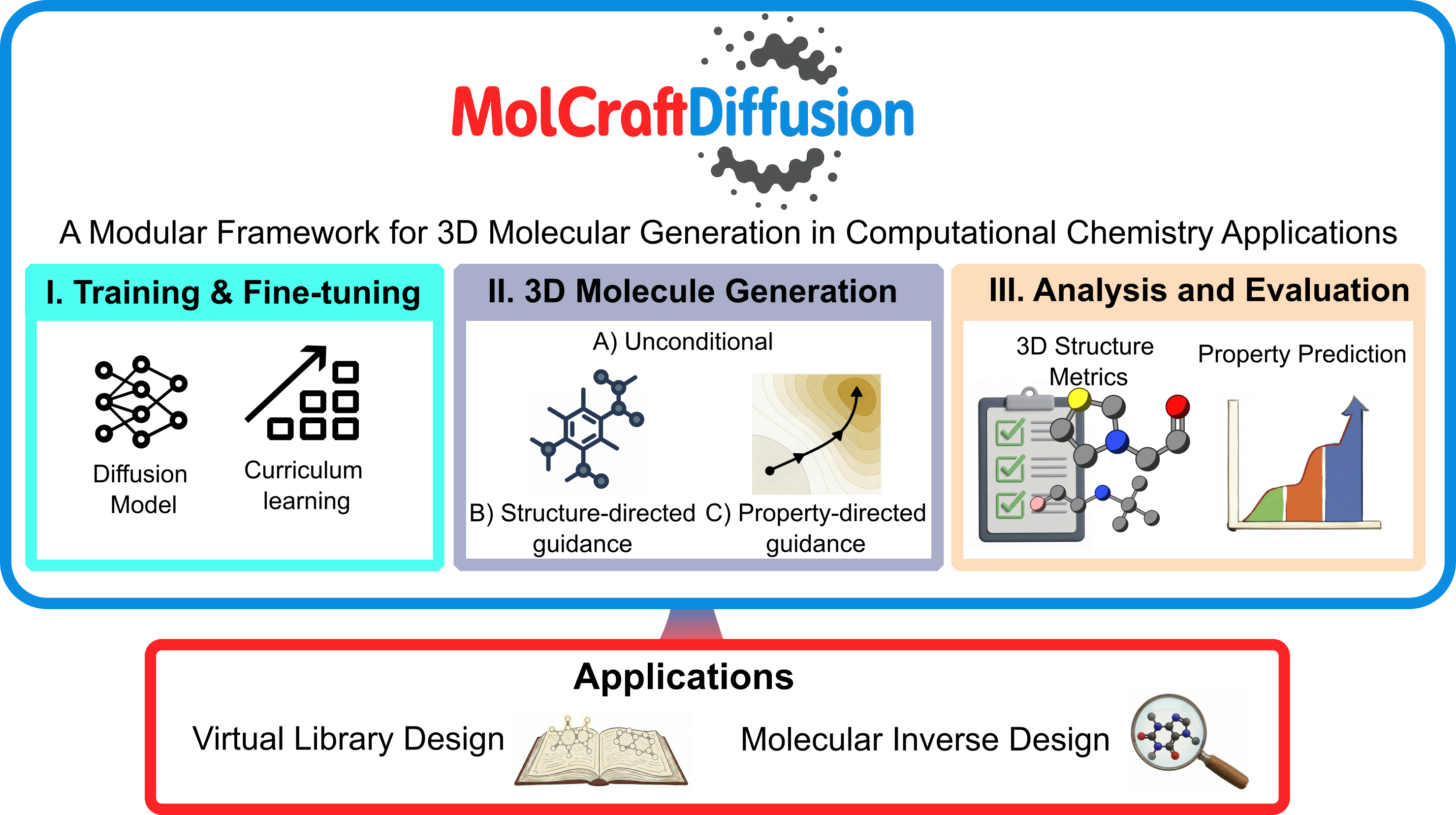
Key Features¶
MolCraftDiffusion is built with modularity at its core, offering an all-in-one, systematic workflow entirely driven by a unified CLI and YAML configuration files.
Data Module — Preprocess, compile, and manage raw
.xyzfiles into unified.db(ASE Database) pipelines, and annotate properties.Training & Fine-Tuning Module — Flexibly train (or fine-tune) diffusion models, property regressors, and time-aware guidance models.
Generation & Guidance Module — Generate 3D molecules using a variety of guidance mechanisms:
Unconditional Generation: Generate 3D molecules without any specific constraints or guidance.
Property-Targeted Guidance: Steer generation toward desired properties using Classifier-Free Guidance (CFG), Gradient Guidance (GG), or a hybrid approach.
Structure-Guided Generation: Perform inpainting (scaffold decoration) and outpainting (fragment extension) with precise 3D geometric constraints.
Analysis & Evaluation Module — Assess the quality of generated 3D molecules. Includes tools for structural validity metrics, xTB geometry optimization, RMSD comparisons, and quantum-chemical property calculation/prediction.
Quick Start¶
# Train a diffusion model
MolCraftDiff train my_config
# Generate molecules
MolCraftDiff generate my_gen_config
# Analyse outputs
MolCraftDiff analyze metrics generated_molecules/
Ready-to-use template configuration files for common workflows are listed in Configuration Templates, with the full packaged Hydra defaults under src/MolecularDiffusion/configs/. Copy the relevant file, fill in your paths, and run:
# Example: unconditional generation with the template
cp docs/cfg_examples/gen_unconditional.yaml my_gen.yaml
# edit my_gen.yaml → set chkpt_directory
MolCraftDiff generate my_gen
Contents¶
Getting Started
Tutorials
- Tutorial 0: Data Preparation & Management
- Tutorial 1: Training a Diffusion Model
- Tutorial 2: Training a Regressor Model
- Tutorial 3: Training a Time-Aware Guidance Model
- Tutorial 4: Fine-Tuning a Diffusion Model
- Tutorial 5: Molecule Generation Overview
- Tutorial 6: Structure-Guided Generation
- Tutorial 7: Property-Directed Generation
- Tutorial 8: Prediction and Evaluation Modes
- Tutorial 9: Analyze Module - 3D Molecular Structure Analysis
- Tutorial 10: Generation Parameter Sweeps
Applications
Workflows
API Reference





