Core Concepts¶
Generators¶
Each generator requires different input and is suited to a different design task.
Generator |
Input required |
Task |
|---|---|---|
Reinvent |
None |
De novo generation atom-by-atom |
LibInvent |
Scaffold SMILES |
Decorate a scaffold with R-groups |
LinkInvent |
Two fragment SMILES |
Find a linker between two fragments |
Mol2Mol |
Reference molecule SMILES |
Generate analogues within a defined similarity radius |
Priors and Agents¶
Prior: a model pre-trained on a large chemical dataset (e.g. ChEMBL) that has learned valid SMILES grammar. It generates molecules without any task-specific bias. Pre-trained priors for all generators are available on Zenodo.
Agent: a copy of the prior whose weights are updated during a run to increase the likelihood of generating high-scoring molecules.
Model Adaptation¶
Two methods can be used independently or in sequence:
Transfer Learning (TL): retrains a prior on a focused SMILES dataset (e.g. known actives for a target), producing an agent biased toward that chemical series.
Reinforcement Learning (RL): iteratively updates the agent using a scoring function as reward signal.
Left: the prior samples broadly across chemical space. Middle: TL narrows the distribution toward a target region. Right: RL concentrates sampling on high-scoring molecules within that region. Taken from Loeffler et al., J. Cheminformatics (2024) under a Creative Commons Attribution 4.0 International License.
Staged Learning / Curriculum Learning¶
RL can be extended into multiple sequential stages. The agent checkpoint from stage N becomes the starting point for stage N+1, each with its own scoring function and termination criterion (max_score, max_steps). This allows gradually increasing objective complexity — e.g. stage 1 filters with fast, cheap components (structural alerts, drug-likeness) while stage 2 introduces expensive ones (docking). Stages can also be used to manually continue a run from any saved checkpoint.
Scoring in RL¶
The scoring function is a weighted combination of components, each computing a property of the generated SMILES (e.g. QED, LogP, TPSA, docking score). Raw values are mapped to [0, 1] via transforms (sigmoid, step, etc.) and aggregated, by default with the geometric mean.
Diversity vs. Exploitation¶
RL naturally drives the agent toward high-scoring regions, but without any check it tends to collapse onto a small set of structurally similar molecules, the same scaffold repeated across the batch (see Figure below). This is undesirable in drug discovery, where we want to explore diverse chemotypes that satisfy the same objective. REINVENT4 addresses this with two mechanisms:
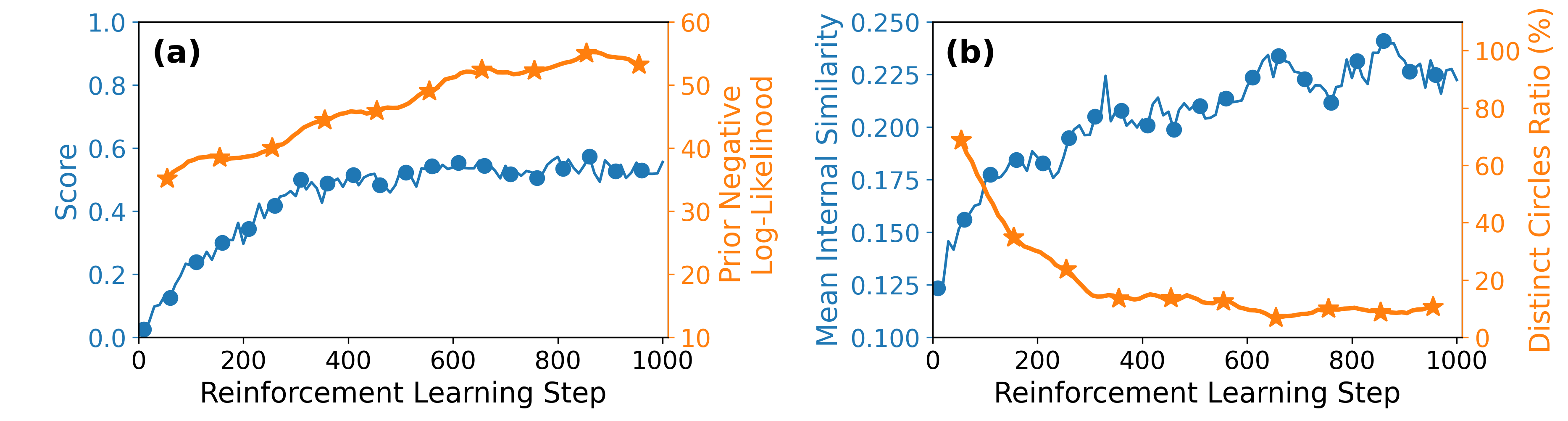 An example of evolution of molecular optimization and diversity during reinforcement learning. Taken from Azzouzi M., Worakul T., et al., Digit. Discov. (2026) under a Creative Commons Attribution 4.0 International License
An example of evolution of molecular optimization and diversity during reinforcement learning. Taken from Azzouzi M., Worakul T., et al., Digit. Discov. (2026) under a Creative Commons Attribution 4.0 International License
REINVENT4 addresses this at two levels:
Diversity Filter: penalises repeated Murcko scaffolds during the run. Molecules are bucketed by scaffold; once a bucket fills, further molecules with that scaffold are penalised. Only molecules above
minscoreenter memory. Types:IdenticalMurckoScaffold(recommended),IdenticalTopologicalScaffold,ScaffoldSimilarity(Tanimoto-based),PenalizeSameSmiles(exact SMILES repetition).Inception (Experience Replay): replays the highest-scoring molecules seen so far alongside the current batch in the loss computation. Useful when high-scoring molecules are rare — prevents the agent from forgetting them between epochs. Memory can be pre-seeded with known actives. Reinvent only.
References:
Olivecrona, M.; Blaschke, T.; Engkvist, O.; Chen, H. Molecular De-Novo Design through Deep Reinforcement Learning. J. Cheminform. 2017, 9 (1), 48. https://doi.org/10.1186/s13321-017-0235-x
Blaschke, T.; Arús-Pous, J.; Chen, H.; Margreitter, C.; Tyrchan, C.; Engkvist, O.; Papadopoulos, K.; Patronov, A. REINVENT 2.0: An AI Tool for De Novo Drug Design. J. Chem. Inf. Model. 2020, 60 (12), 5918–5922. https://doi.org/10.1021/acs.jcim.0c00915
Fialková, V.; Zhao, J.; Papadopoulos, K.; Engkvist, O.; Bjerrum, E. J.; Kogej, T.; Patronov, A. LibINVENT: Reaction-Based Generative Scaffold Decoration for in Silico Library Design. J. Chem. Inf. Model. 2022, 62 (9), 2046–2063. https://doi.org/10.1021/acs.jcim.1c00469
Guo, J.; Fialková, V.; Arango, J. D.; Margreitter, C.; Janet, J. P.; Papadopoulos, K.; Engkvist, O.; Patronov, A. Improving de Novo Molecular Design with Curriculum Learning. Nat. Mach. Intell. 2022, 4 (6), 555–563. https://doi.org/10.1038/s42256-022-00494-4
Guo, J.; Knuth, F.; Margreitter, C.; Janet, J. P.; Papadopoulos, K.; Engkvist, O.; Patronov, A. Link-INVENT: Generative Linker Design with Reinforcement Learning. Digit. Discov. 2023, 2 (2), 392–408. https://doi.org/10.1039/D2DD00115B
Loeffler, H. H.; He, J.; Tibo, A.; Janet, J. P.; Voronov, A.; Mervin, L. H.; Engkvist, O. Reinvent 4: Modern AI-Driven Generative Molecule Design. J. Cheminform. 2024, 16 (1), 20. https://doi.org/10.1186/s13321-024-00812-5
Guo, J.; Schwaller, P. Augmented Memory: Sample-Efficient Generative Molecular Design with Reinforcement Learning. JACS Au 2024, 4 (6), 2160–2172. https://doi.org/10.1021/jacsau.4c00066